When we learned about making our won tools to keep us healthy and detox from all the metals, nano-stuff and poisons like 15 years ago, many of us produced his different tools. I mainly made the tools with magnets and kept away from the electrical wires etc. But times have changed and we have been bombarded with all kind of poisonous stuff. Here are some articles from Tony Pantalleresco, you find them, by surfing more “deeply” all over the internet. It takes some time, as many have disconnected links (on purpose!).
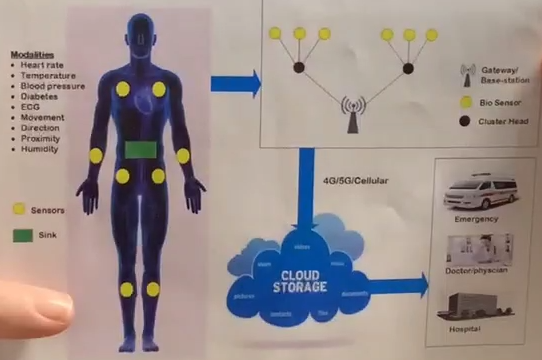
Having worked for over 30 years with our various “body parts”, like the physical body, the emotional body, the mental body, the lightbody and our aura or biofield that surrounds us and is interconnected with our neurons, many energy activities, there is still for most a part missing. The new technology we have today, that is bombarding us day and night, connecting to our bio sensors in our bodies forces us to learn to neutralize them and to to learn how to take our DNA radiation for moments back into our spinal core. Tony Pantalleresco and Sabrina from Psinergy can help understand connection many dots that were not seen till today.
How to make your own anti-nano bucket with Tony: Anti-Nano Station
Article 2015:
Viruses can be made to churn out high-tech nanomaterials
Viruses subvert their hosts to pump out masses of new viruses. In an unusual twist, an MIT researcher reports in the May 3 issue of Science that she used genetically engineered viruses that are noninfectious to humans to mass produce tiny materials for next-generation optical, electronic and magnetic devices.–“We’ve been looking at using genetic tools to grow semiconductor materials,” said author Angela M. Belcher, associate professor of materials science and engineering and biological engineering. “In this case, we took advantage of the viruses’ genetic makeup and physical shape to not only grow the material but also to help them assemble themselves into liquid crystal structures that are several centimeters long.”–Belcher and colleagues at the University of Texas at Austin are interested in using the processes by which nature makes materials to design new biological-electronic hybrid materials that could be used to assemble electronic materials at the nanoscale. Her research brings together inorganic chemistry, materials chemistry, biochemistry, molecular biology and electrical engineering. She will join the MIT Department of Material Science and Engineering and the Biological Engineering Division of the School of Engineering in September.–Belcher’s approach is to use systems such as viruses that evolved over millions of years to work perfectly at the nanoscale, but to convince the viruses to work on technologically important materials. Belcher’s research team can evolve the viruses to work on the materials of interest over a period of months.–Building self-assembling and defect-free two- and three-dimensional materials on the nanometer scale is essential for the construction of new devices for optics and electronics. Researchers have been looking at ways to use organic materials to organize molecules of inorganic materials on the nanoscale. Fabricating viral films, Belcher said, may provide new pathways for organizing molecules to help create electronic, optical and magnetic materials.
“We showed that engineered viruses can recognize specific semiconductor surfaces, and these recognition properties can be used to organize molecules in inorganic nanocrystals, forming ordered arrays,” she said. “In this system, we can easily modulate the length of the bacteriophage (the type of virus) and the type of inorganic materials through genetic modification and selection. One can easily modulate and align different types of inorganic nanocrystals in 3D layered structures.”
This work is supported by the Army Research Office, the National Science Foundation and the Welch Foundation.
****************************************************************************
Bio-inspired synthesis of metal nanomaterials and applications
Jiale Huang† a, Liqin Lin† b, Daohua Sun a, Huimei Chen c, Dapeng Yang *b and Qingbiao Li *abc
aDepartment of Chemical and Biochemical Engineering, College of Chemistry and Chemical Engineering, and National Laboratory for Green Chemical Productions of Alcohols, Ethers, and Esters, Xiamen University, Xiamen, P. R. China. E-mail: kelqb@xmu.edu.cn; Fax: +86 592 2184822; Tel: +86 592 2184822
bCollege of Chemistry and Life Science, Quanzhou Normal University, Quanzhou, P. R. China. E-mail: yangdp@qztc.edu.cn
cEnvironmental Science Research Center, College of the Environment & Ecology, Xiamen University, Xiamen 361005, P. R. China
Received 10th February 2015
First published on the web 17th June 2015
2.
Nano-enabled synthetic biology
Mitchel J Doktycza,1,2 and Michael L Simpsonb,1,3,4
Author information ► Article notes ► Copyright and License information ►
This article has been cited by other articles in PMC.
Abstract
Biological systems display a functional diversity, density and efficiency that make them a paradigm for synthetic systems. In natural systems, the cell is the elemental unit and efforts to emulate cells, their components, and organization have relied primarily on the use of bioorganic materials. Impressive advances have been made towards assembling simple genetic systems within cellular scale containers. These biological system assembly efforts are particularly instructive, as we gain command over the directed synthesis and assembly of synthetic nanoscale structures. Advances in nanoscale fabrication, assembly, and characterization are providing the tools and materials for characterizing and emulating the smallest scale features of biology. Further, they are revealing unique physical properties that emerge at the nanoscale. Realizing these properties in useful ways will require attention to the assembly of these nanoscale components. Attention to systems biology principles can lead to the practical development of nanoscale technologies with possible realization of synthetic systems with cell-like complexity. In turn, useful tools for interpreting biological complexity and for interfacing to biological processes will result.
Keywords: biological systems, carbon nanofibers, cell mimic, nanotechnology, synthetic biology, synthetic cells
Introduction
Understanding the organizing principles of complex systems presents a significant challenge. Whether in the synthetic or biological domain, there is a growing awareness that design, the reiterative process of creating function through the intentional interconnection of components, is an indispensable tool for unraveling complexity. Whereas modeling, simulation, and experimental analyses have a tendency to focus attention on the details of individual elements, design requires grappling with the trade-offs and compromises needed to enable system function. Along these lines, synthetic biology efforts follow a strategy of constructing deliberately simplified systems to comprehend molecular and cellular regulatory processes from the bottom up (Hasty et al, 2002; Sprinzak and Elowitz, 2005; Andrianantoandro et al, 2006; Guido et al, 2006). Similarly, efforts towards constructing minimal cells either add, subtract or manipulate components to realize simple systems with desired capabilities (Forster and Church, 2006; Luisi et al, 2006). In both cases, iterative design is a fundamental aspect of the approach and represents a major step towards true bottom-up construction of biological complexity. However, these approaches still depend on a platform of an existing cellular environment or the use of biomolecules (nucleic acids, proteins, lipids) to jump start cellular function. Thus, the question remains, what could be learned from a true bottom-up effort to reconstitute cell-like complexity?
Can the deliberate design and assembly of synthetic components lead to systems with cell-like characteristics? The large discrepancy between the functional density (i.e., the number of components or interconnection of components per unit volume) of cells and engineered systems highlights the inherent challenges posed by such a question. A simple example compares Escherichia coli (∼2-μm2 cross-sectional area) with an equivalent area on a silicon-integrated circuit (Simpson et al, 2001). The E. coli cell has an ∼4.6-million base-pair chromosome (the equivalent of a 9.2 megabit memory) that codes for as many as 4300 different polypeptides under the inducible control of several hundred different promoters, whereas the same space on a silicon chip could provide only a very small fraction of this memory or a few simple logic gates. Clearly, the operational scale of biological systems is significantly smaller than that of conventionally engineered systems. Beyond just density alone, it is also the drastically different approach to component assembly, interfacing, and organization that differentiates the biological from the synthetic nanoscale system. In the biological substrate, dynamic systems exploit weak interactions, arranged to provide desired specificity, and take place in a fluid environment. These features lead from simply high spatial density to high functional density and the realization of robust, adaptable systems.
As nanoscience and technology advance, the opportunity to match the scale of biological system components becomes feasible. As a first step, nanotechnology presents the ability to directly interface to the working levels of biology, leading to the emergence of new approaches to therapy and diagnostics. Additionally, the emulation of biological design principles using synthetic components becomes feasible. Potentially, as systems of such elements approach biological-scale functional density, they can begin to assume cell-like characteristics including: (1) construction from an inhomogeneous mixture of materials with different properties, modes and strengths of interactions, and relative abundances; (2) the encoding of information within small populations (e.g., biomolecules or electrons); (3) function emerging from an environment with large stochastic fluctuations (a consequence of (2)); and (4) the efficient transduction of information, energy, and materials that emanates from the molecular scale. It is an intriguing possibility that, as our ability to control the synthesis and direct the assembly of synthetic nanoscale elements increases, we may attempt the bottom-up design and construction of nanosystems with cell-like complexity and capabilities. In turn, the design of such systems will lead to an enhanced ability to understand and interface to biological systems.
The intersection of nanoscale science and technology with biology has figured prominently in even the early stages of envisioning nanoscience research directions and goals (Roco, 2003). In many ways, the biological cell represents an ideal paradigm for nanoscale systems. Being the fundamental unit of biological systems, their function can be extremely diverse, yet uses only a finite, common set of building blocks. Cells operate under a wide range of environmental conditions with efficiencies unmatched by artificial systems. They can be highly specialized and carry out tens of thousands of chemical reactions in parallel. The dimensional characteristics of cells are well conserved and undoubtedly critical for system function (Welch, 1992; Hess and Mikhailov, 1994; Hochachka, 1999; Misteli, 2001; Harold, 2005). Short distances (nm–μm) enable intra- and intercellular communication using simple diffusion-based mechanisms. Also, the small fluid volume of a cell allows for small fluctuations in numbers of specific molecules to result in dramatic changes in cellular state. Higher order, nanoscale structuring (Welch, 1992; Hochachka, 1999) and excluded volume effects (Hall and Minton, 2003) are also known to be critical to cellular function. In fact, with regard to heredity, the spatial definition of the cell may be as important as the genetic material (Harold, 2005).
Here, we consider the potential for a nano-enabled synthetic biology that may be derived from the confluence of systems biology and nanoscale science and technology. At this confluence, systems biology provides knowledge of the chemical components that comprise the cell and the spatial and temporal interplay between these components. Initial efforts to mimic cells have followed a path of using soft materials that are similar or identical to cellular materials. However, the continued progress of nanoscale science and technology provides hope that many cellular attributes may be transferred to artificial systems through the control of the synthesis and assembly of hard nanoscale materials at the multiple size scales important to cellular function. In the process, advanced tools for understanding basic questions regarding biological function will be provided. Such developments could benefit both technology and science. Cell-like complexity in nanoscale systems may lead to significantly higher levels of function, whereas also forming an experimental system that would allow a much better examination of cellular organizational principles. Here, we highlight efforts to mimic cell-like systems and the emerging tools of nanoscience that may enable an even more synthetic biology.
Mimicking cellular systems
The general concept of mimicking cells dates back several decades with the initiation of efforts to make effective blood substitutes (Chang, 2004). More recently, multiple efforts have evolved and are focused on engineering molecular systems and mimicking functional aspects of cells. Additionally, synthetic cell efforts are increasingly integrating synthetic materials and nanotechnologies. A common feature of cell mimic pursuits is containment of a small, aqueous volume. The ability to contain small volumes of liquid (pico– to nanoliter) is a critical aspect of biological cells and enabling for the creation of synthetic cell-like systems. Small volume containers obviate the need for mixing and establish local conditions that are favorable for protein function. Small volumes reduce the number of molecules needed for carrying out a function. Therefore, they are ideal for studying, or exploiting, reactions that involve single molecules. Further, small volume containers can be used for understanding molecular reaction systems and self organization at the cellular scale (Hess and Mikhailov, 1995; Marijuan, 1995; Chiu et al, 1999; Misteli, 2001; Long et al, 2005; Pielak, 2005). They are also valuable for studies related to understanding questions involving the origin of life (Deamer, 2005). On the applied side, miniaturization of the reaction volume can lead to the creation of massively parallel analytical systems (Wolcke and Ullmann, 2001; Heller, 2002; Khandurina and Guttman, 2002), whereas the in vitro aspects of the technology allow the use of physical conditions or the synthesis of products that may be toxic to natural cells. New approaches to high throughput screening, chemical sensing, and drug delivery are being enabled. The incorporation of synthetic nanomaterials will be key to realizing these diverse applications. The containing ‘membrane’ is a distinguishing feature of present approaches to mimic functional aspects of cells (Figure 1). Natural membrane components, synthetic polymers, emulsion systems, and microfabricated structures are being considered. An overview of these different systems is described below.
Overview of a genetic-based synthetic cell. The membrane of synthetic cells can be created from a variety of materials, including natural membrane components, synthetic polymers, and micro- or nano-fabricated materials. Alternatively, a water-in-oil emulsion …
Vesicle-based systems
The most widely studied biomimetic containment systems are based on vesicles prepared from amphiphilic molecules. These self-assembling structures can be formed from lipids, creating liposomes, or from synthetic molecules such as block copolymers, which are often referred to as polymersomes (Vriezema et al, 2005). They are also considered to be ideal biomimetic nanoscale reaction containers (Karlsson et al, 2004). Liposomes have long been used to encapsulate enzymes and can be prepared using a variety of techniques (Walde and Ichikawa, 2001). Their potential application as delivery vehicles for therapeutics has garnered much attention. Liposomes can protect enzymes from degradation, effect slow release of a reagent, or contain chemical reactions. For example, enzymes entrapped in the interior of the liposome can be used for diagnostic applications ((Yoshimoto et al, 2003). Further, multi-component systems have been designed that allow for targeting and stimuli-dependent release of encapsulated reagents (Guo and Szoka, 2003).
Gene-based reactions systems are also being developed. Reactions involving nucleic acids and polymerases have been described (Walde and Ichikawa, 2001; Monnard, 2003). Further, simple genetic constructs, involving a promoter and gene sequence, and cell-free extract (Spirin et al, 1988; Shimizu et al, 2001) can be pooled in liposomes to produce the corresponding protein. The expression of green fluorescent protein) enables easy assessment of the reaction (Yu et al, 2001; Oberholzer and Luisi, 2002; Nomura et al, 2003; Ishikawa et al, 2004; Sunami et al, 2006). More complex reactions have also been demonstrated. For example, Ishikawa et al (2004) demonstrated a two-stage genetic network, where the protein product of the first stage is necessary for driving protein synthesis of the second stage. Other multi-stage reaction systems have been described leading to the possibility of constructing cell-free genetic circuits (Noireaux et al, 2003; Noireaux and Libchaber, 2004).–The use of natural lipids facilitates biocompatibility, including the use of membrane proteins to facilitate material exchange with the enclosed volume. However, the long-term stability of these structures can be problematic. Nanotechnologies can effectively address this shortcoming. Related efforts have investigated the use of synthetic polymers to create polymersomes. Block copolymers are finding multiple applications in nanotechnology. These polymers are composed of at least two parts of differing solubility and can self assemble into a variety of structures (Forster and Antonietti, 1998; Klok and Lecommandoux, 2001; Park et al, 2003). They can also be formed into vesicles. Vesicles with a broad range of chemistries and physical properties that are based on the choice of polymer type, block ratio, and molecular weight, can be constructed (Discher and Eisenberg, 2002). As with liposomes, applications in chemical sensing, reagent delivery and reaction containment are pursued. For example, enzyme activity can be preserved when encapsulated within polymersomes and such systems can be used for sensing and can be made stimulus responsive (Napoli et al, 2004a, 2004b). Facilitating the use of polymersomes for incorporation into biological systems is the discovery that natural membrane spanning proteins can incorporate into block copolymer shells. Nardin et al (2001) demonstrated that the E. coli porin protein OmpF can form a stable protein/polymer hybrid membrane and act as a size selective filter. In this case, protein incorporation occurs even though the polymer membrane is two- to threefold thicker than a conventional lipid bilayer. Subsequent work has demonstrated the ability to use other block copolymer systems and incorporate other proteins (Ho et al, 2005; Ranquin et al, 2005). For example, Ho et al (2005) have incorporated the energy-transducing membrane proteins bacteriorhodopsin and cytochrome c oxidase into block copolymer vesicles. This system was shown to generate transmembrane pH gradients and highlights the potential use of hybrid nanosystems for harnessing the energy conversion processes of natural systems.
Emulsion-defined systems
Small volume reaction containers can also be created by water in oil (w/o) emulsions (Tawfik and Griffiths, 1998; Griffiths and Tawfik, 2000, 2003; Ghadessy et al, 2001; Pietrini and Luisi, 2004). Enclosed, femtoliter scale volumes can be defined through simple shaking, stirring or extrusion of a mixture containing an aqueous solution, oil and appropriate surfactants to stabilize the emulsion. The typical size of the water droplets is on the order of a few microns, comparable to that of a microbial cell. Even smaller droplets can be prepared using ultrasonication (Musyanovych et al, 2005). The simplicity and ability to create ∼1010 containers/ml volume has enabled a variety of applications. For example, complete genomic libraries can be created, amplified and characterized using w/o techniques (Margulies et al, 2005; Shendure et al, 2005). Alternatively, genetic variation within individual alleles can be quantified (Dressman et al, 2003). In these applications, the DNA is diluted such that, on average, a single template is contained within an aqueous droplet. These individual DNA strands can then be amplified by the polymerase chain reaction in separate aqueous volumes within a single tube.
A notable feature of the w/o emulsion technique is the ability to link genotype with phenotype in a small volume reactor. An in vitro compartmentalization system has been described that is useful for high-throughput screening and selection (Aharoni et al, 2005b). In this approach, a gene sequence, along with the appropriate reagents for transcription and translation, is contained within the aqueous compartment. As a large number of compartments can be created and tested simultaneously, entire libraries of genetic variants can be assessed. This enables ‘directed evolution’ of protein function by selection for the appropriate activity. For example, DNA polymerases (Ghadessy et al, 2001) and methyl transferases (Tawfik and Griffiths, 1998; Lee et al, 2002) have been selected for by simply breaking the emulsion and identifying the remaining gene sequences. Other approaches exploit a physical connection between the gene and its product (Doi and Yanagawa, 1999; Sepp et al, 2002; Griffiths and Tawfik, 2003) or novel approaches (Aharoni et al, 2005a; Mastrobattista et al, 2005) to allow for subsequent sorting.
Enhancing the ability to transport reagents into and out of w/o emulsion-based reaction vessels is still under investigation. In general, reaction extent inside the vessel is limited by the availability of precursor reagents, as transport within the oil phase is unlikely. Some reagent exchange is believed to take place upon contact between individual compartments (Ghadessy et al, 2001). Fusion of compartments is also a potential mechanism for exchanging or feeding reagents to reaction vessels (Bernath et al, 2004; Pietrini and Luisi, 2004). A related approach exploits microfluidics technology for creating aqueous droplets of defined size and composition (Dittrich et al, 2005). Specific reagents can be mixed and compartmentalized by the merging of microfluidic flow streams. This approach allows facile manipulation of the droplet and can be combined with sensitive detection techniques.
Nanomaterials: from individual elements to cell-like complexity
As described above, most efforts to mimic cells have relied on the self-assembly properties of organic materials. However, many applications have benefited from small scale and high parallelism afforded by advanced microfabrication techniques, and these techniques have become better integrated with biological materials to enable greater functionality. One use of these fabrication techniques is for creating robust reaction containers of defined volume and contents. Various etching, drilling, embossing or molding techniques can be used to create containers of a range of sizes (zl–μl). The integration of such structures with fluids and biological materials is enabling multiple applications. For example, small volume reaction containers are being considered for high-throughput screening, (Grosvenor et al, 2000; Angenendt et al, 2005), single molecule enzymology (Rondelez et al, 2005; Rissin and Walt, 2006), and analyses of single cells (Cooper, 1999; Johannessen et al, 2002). These reaction containers are also being developed for the cell-free synthesis of proteins (Nojima et al, 2000; Tabuchi et al, 2002; Yamamoto et al, 2002; Angenendt et al, 2004; Kinpara et al, 2004; Mei et al, 2005). In addition to creating a new approach to the parallel production of various proteins, these structures are permitting novel functional assays (Angenendt et al, 2005; Mei et al, 2005). The use of microfabricated structures allows for the controllable exchange and mixing of reagents (Nojima et al, 2000; Wang et al, 2005) and the integration of sensitive techniques for sampling and analysis of reaction products.
However, the use of microfabrication technology to achieve or interface to cell-like complexity is ultimately limited by the shortcomings of top-down synthesis processes that require layer-by-layer definition of structure through a very well-controlled series of deposition, lithography, and etching steps. Instead, synthetic systems must exploit characteristics similar to natural components, and nanoscale materials are especially suited to this challenge as they reside on the same size scale as the components of biological processes, whereas exhibiting electrical, magnetic, optical, thermal, and chemical properties conducive to the construction of complex networks of functional parts. By definition, nanoscale materials have a limited extent (nominally defined as less than 100 nm) in at least one of the three spatial dimensions. Zero-dimensional (0-D) materials include semiconductor quantum dots (QD), colloidal metal particles, and atomic or molecular clusters that are confined in all three spatial dimensions; 1-D structures (which we refer to collectively as nanowires) are confined in two spatial dimensions; and 2-D structures (thin films such as silicon nitride or lipid bilayer membranes) are confined in only one spatial dimension (Figure 2).
Collage of synthetic nanoscale materials. (A) 0-d nanoscale material shown in a Z-STEM image of 655 QdotsTM (CdSe/CdS core/shell, Quantum Dot Corporation; image courtesy of Professor S Rosenthal, Vanderbilt University (adapted with permission from McBride …
Much of the early effort in nanoscience has focused on the synthesis and characterization of individual or homogeneous arrays of nanoscale elements, and numerous techniques have been developed for the synthesis of a variety of nanoscale materials: QDs composed of periodic groups II–VI (e.g., CdSe) or III–V (e.g., InP) materials are synthesized by injecting liquid precursors into hot (300°C) coordinating organic solvent (Murray et al, 1993; Peng et al, 1998); semiconductor 1-d nanowires can be grown in a vapor-liquid-solid process (Wagner and Ellis, 1964) in which a liquid metal cluster or catalyst acts as the energetically favored site for absorption of gas-phase reactants; and carbon nanowires, which include carbon nanotubes (Ajayan and Ebbesen, 1997) and carbon nanofibers (CNFs) (Rodriguez, 1993; Melechko et al, 2005), are synthesized in numerous processes including laser-vaporization (Kroto et al, 1985), arc discharge (Kratschmer et al, 1990; Iijima, 1991), catalytic chemical vapor deposition, and catalytic plasma-enhanced chemical vapor deposition (C-PECVD).
Although these materials synthesis efforts have been foundational for nanoscale science, taken alone they do not provide the means to construct or interface to cell-like complexity. It is the collective behaviors of interacting nanoscale components, where scale and complexity lead to ‘entirely new properties’ (Anderson, 1972). The question then is not only of the synthesis of nanomaterials, but also that of how these materials should be organized into ensembles that exhibit new levels of functionality. Addressing this question shifts the focus from synthesis of individual elements to the controlled synthesis and directed assembly of systems of nanoscale components that are capable of assuming cell-like organization. By controlled synthesis, we mean a process of mass nanostructure growth, where the pertinent attributes (location, size, orientation, composition, electrical, mechanical, and thermal properties, etc.) of the individual elements can be selected a priori by the choice of the growth conditions or the preparation of the growth substrate. Much like biological materials (e.g., silica biomineralization (Morse, 1999; Hildebrand, 2003; Hildebrand et al, 2006)), or embryogenesis (Carroll et al, 2004)), directed assembly in this context may more appropriately be thought of as hierarchical assembly, as each stage in the process forms the template for the next layer of added complexity. We illustrate these concepts in a synthetic system using the example of CNFs below. In this example, we emphasize the use of self-organization, hierarchical assembly, and the emergence of functional order from stochastic processes.
Controlled synthesis and hierarchical assembly
Carbon nanofibers are grown in a PE-CVD process from metal catalyst materials supported on substrates of various types (Melechko et al, 2005). The complex plasma environment can be manipulated to produce changes in nanofiber aspect ratio, diameter, orientation, shape, and chemical composition (Merkulov et al, 2001, 2002a, 2002b, 2002c; Melechko et al, 2002). Likewise, CNF morphology, crystalline structure, and composition can be varied through manipulation of the growth substrate (Klein et al, 2005; Fowlkes et al, 2006b). For example, the patterning of the catalyst material allows the selection of either randomly spaced forests or deterministically placed isolated fibers (Figure 3); the selection of the plasma source gases control nanofiber composition; and nanofiber shape can be controlled by selection of the catalyst material crystallographic orientation (Fowlkes et al, 2006b). The forest morphology is particularly interesting, as it is the result of a self-organization process initiated by the plasma-induced fragmentation and reordering of the initial microscale catalyst pattern followed by nanofiber growth from each of the nanoparticles. Although the placement of individual nanofibers exhibits a high degree of stochasticity, the distributions of interfiber spacing and fiber size are strongly influenced by the choice of catalyst material, thickness, and crystallographic orientation.
Micrographs of carbon nanofibers. (A) Vertically aligned carbon nanofibers can be prepared from a continuous catalyst stripe yielding an array of CNFs that are randomly arranged. (B) Individual CNFs can be precisely positioned using electron beam lithography …
A concurrent self-organization is the emergence of nanoscale pores that form between the randomly spaced nanofibers. These structures can act as passive membrane mimics in microfluidic devices (Zhang et al, 2002; Fletcher et al, 2004). Whereas the pores are relatively large (e.g., 150–250 nm) in as grown forests, hierarchical processing that adds to the diameters of the nanofibers (e.g., conformal SiO2 coating or electroplating of conductive polymers) leads to the formation of planar or 3-D pore networks (Fowlkes et al, 2005). Within these forest structures, diffusive transport is a strong function of the excluded volume (i.e., space taken up by the nanofibers) and the placement of nanofibers. The stochastic nature of the interactions between diffusing molecules and nanofibers leads to regions of anomalous diffusion (i.e., time-variant diffusion coefficient), whereas a large excluded volume leads to significantly reduced molecular mobility (Fowlkes et al, 2006a). The net effect is that the diffusive transport properties of these membrane mimics may be controlled through self-organization of stochastic nanofiber forests and hierarchical processing to define the structure of a nanoporous network.
These membrane structures can be patterned in arbitrary shapes and used for creating cell mimic structures (Fletcher et al, 2004; Fowlkes et al, 2005). As shown in Figure 4, advanced, multiscale (nano to macro) fabrication techniques allow for the integration of nanomaterials, fluids, and biological reagents to create structures that mimic functional characteristics of a cell. In this example, the enclosed volume can be tuned to closely match those of a natural cell. Such a container would be useful for experimentally characterizing reaction systems and material organization in a contained fluid environment. Further, this structure allows for controlled transport between the contained volume and the local environment through design of the membrane properties.
Fabrication of a cell mimic array. The device is created using (A) contact photolithography and ICP-RIE etching of silicon to define a fluidic channel approximately 10 μm in depth. (B, C) Metal lift-off of Ni catalyst is followed by PECVD growth …
These structures can advance from being simple, passive structures that control transport based on physical size to more sophisticated, active nanostructures. For example, additional control of the membrane properties is possible through the application of chemical coatings (Fowlkes et al, 2005; Fletcher et al, 2006; McKnight et al, 2006). Such coatings can bestow chemical specificity in addition to size-selective transport. Polymeric coatings on the CNFs can be exploited to create active interfaces. Polypyrrole can be selectively patterned onto CNF-based electrodes (Chen et al, 2001; Nguyen-Vu et al, 2006; Fletcher et al, 2007). Such coatings can be reversibly actuated to expand or contract with the application of an electrical signal (Smela, 2003). Other polymers are responsive to chemical or physical stimuli (Gil and Hudson, 2004; Yoshida et al, 2006).
Realizing a nano-enabled synthetic biology
Nano-enabled synthetic biology is in its early stages. For biological systems, functionality at any scale begins at the cellular level. The efforts described above illustrate the different approaches involved in attempting the bottom-up construction of simple cellular-like structures. Nanotechnologies are becoming increasingly involved. Nanoscale science has delivered the ability to synthesize a variety of nanoscale components that provide the means to dramatically increase the density of elements. Controlled synthesis and hierarchical assembly allow these components to mimic passive, and even active, cell-like behaviors. As described, carbon-based nanostructures and block co-polymers match many of the design requirements of synthetic membranes. Future efforts will likely see the integration of other nanomaterials for potentially transducing energy, conveying signals, or controlling the arrangement of biological and synthetic structures.
Understanding and improving the interface between natural and synthetic structures represents a key next step for nano-enabled synthetic biology. Related challenges in controlling synthesis across the multiple length scales relevant to biology and in the development of tools that are useful in characterizing interactions at these scales also need continued attention. Effective emulation, interpretation, or control of biomolecular events will depend on this interface. As stated at the beginning of this article, it is both scale and complexity (interconnectivity of the elements) that lead to higher levels of functionality, and further progress hinges on increases in the latter. Perhaps paradoxically, increasing complexity requires a renewed, albeit redirected, focus on synthesis. Nanoscale materials have been pursued with an eye on unique properties that emerge usually due to electron confinement, the increased ratio of surface area to volume, or the physical properties that result from precise molecular arrangements. Increased attention towards control of surface properties to allow site-specific functionalization of nanomaterials will be required. The binding affinity between natural and synthetic structures will need to be carefully prescribed. Nanoelements will need to be multifunctional, possibly requiring a mix of soft/hard material functionality and hybrid synthesis techniques that. Synthetic nanostructures will need to become active participants in the feedback mechanisms of biological networks for effective interfacing.
Ultimately, the development of nanotechnology-based tools will enable hybrid systems that will substantially enhance the synthetic biology toolbox. Practical biomedical devices will also result. Of course, interfacing to biological systems is not a requirement for systems composed of synthetic nanoscale components. As nanostructured materials take on the characteristics of biological materials, synthetic systems of high functional density and cell-like complexity may also be realized. Learning how to assemble these components into functional networks will require a close coupling with systems biology efforts. Such bio-inspired nanomaterial systems would not be restricted to operation in aqueous environments or a narrow range of physical conditions. Considering the diversity observed in biology and the commonality of their system architecture, even simple synthetic systems have the potential for addressing multiple applications.
Acknowledgments
We gratefully acknowledge the staff of the Molecular-Scale Engineering and Nanoscale Technologies and the Biological and Nanoscale Systems research groups for contributions and critiques of the manuscript. Our efforts are supported by NIH Grant EB000657, the Center for Nanophase Materials Sciences, which is sponsored at Oak Ridge National Laboratory by the Division of Scientific User Facilities, US Department of Energy and the DOE Office of Science. This work was performed at the Oak Ridge National Laboratory, managed by UT-Battelle, LLC, for the US DOE under Contract no. DE-AC05-00OR22725. This manuscript has been authorized by a contractor of the US Government under contract DE-AC05-00OR22725. Accordingly, the US Government retains a nonexclusive, royalty-free license to publish or reproduce the published form of this contribution, or allow others to do so, for US Government purposes.
References
- Aharoni A, Amitai G, Bernath K, Magdassi S, Tawfik DS (2005a) High-throughput screening of enzyme libraries: Thiolactonases evolved by fluorescence-activated sorting of single cells in emulsion compartments. Chem Biol 12: 1281–1289 [PubMed]
- Aharoni A, Griffiths AD, Tawfik DS (2005b) High-throughput screens and selections of enzyme-encoding genes. Curr Opin Chem Biol 9: 210–216 [PubMed]
- Ajayan PM, Ebbesen TW (1997) Nanometre-size tubes of carbon. Rep Prog Phys 60: 1025–1062
- Anderson PW (1972) More is different—broken symmetry and nature of hierarchical structure of science. Science 177: 393–396 [PubMed]
- Andrianantoandro E, Basu S, Karig DK, Weiss R (2006) Synthetic biology: new engineering rules for an emerging discipline. Mol Syst Biol 2: 2006.002816738572 [PMC free article] [PubMed]
- Angenendt P, Lehrach H, Kreutzberger J, Glokler J (2005) Subnanoliter enzymatic assays on microarrays. Proteomics 5: 420–425 [PubMed]
- Angenendt P, Nyarsik L, Szaflarski W, Glokler J, Nierhaus KH, Lehrach H, Cahill DJ, Lueking A (2004) Cell-free protein expression and functional assay in nanowell chip format. Anal Chem 76: 1844–1849 [PubMed]
- Bernath K, Hai MT, Mastrobattista E, Griffiths AD, Magdassi S, Tawfik DS (2004) In vitro compartmentalization by double emulsions: sorting and gene enrichment by fluorescence activated cell sorting. Anal Biochem 325: 151–157 [PubMed]
- Carroll S, Grenier J, Weatherbee S (2004) From DNA to Diversity. Massachusetts: Blackwell Science Malden
- Chang TM (2004) Artificial cells for cell and organ replacements. Artif Organs 28: 265–270 [PubMed]
- Chen JH, Huang ZP, Wang DZ, Yang SX, Wen JG, Ren ZR (2001) Electrochemical synthesis of polypyrrole/carbon nanotube nanoscale composites using well-aligned carbon nanotube arrays. Appl Phys A-Mater Sci Process 73: 129–131
- Chiu DT, Wilson CF, Ryttsen F, Stromberg A, Farre C, Karlsson A, Nordholm S, Gaggar A, Modi BP, Moscho A, Garza-Lopez RA, Orwar O, Zare RN (1999) Chemical transformations in individual ultrasmall biomimetic containers. Science 283: 1892–1895 [PubMed]
- Cooper JM (1999) Towards electronic Petri dishes and picolitre-scale single-cell technologies. Trends Biotechnol 17: 226–230 [PubMed]
- Deamer D (2005) A giant step towards artificial life? Trends Biotechnol 23: 336–338 [PubMed]
- Discher DE, Eisenberg A (2002) Polymer vesicles. Science 297: 967–973 [PubMed]
- Dittrich PS, Jahnz M, Schwille P (2005) A new embedded process for compartmentalized cell-free protein expression and on-line detection in microfluidic devices. Chembiochem 6: 811–814 [PubMed]
- Doi N, Yanagawa H (1999) STABLE: protein–DNA fusion system for screening of combinatorial protein libraries in vitro. Febs Lett 457: 227–230 [PubMed]
- Dressman D, Yan H, Traverso G, Kinzler KW, Vogelstein B (2003) Transforming single DNA molecules into fluorescent magnetic particles for detection and enumeration of genetic variations. Proc Natl Acad Sci USA 100: 8817–8822 [PMC free article] [PubMed]
- Fletcher BL, Hullander ED, Melechko AV, McKnight TE, Klein KL, Hensley DK, Morrell JL, Simpson ML, Doktycz MJ (2004) Microarrays of biomimetic cells formed by the controlled synthesis of carbon nanofiber membranes. Nano Lett 4: 1809–1814
- Fletcher BL, McKnight TE, Fowlkes JD, Allison DP, Simpson ML, Doktycz MJ (2007) Controlling the dimensions of carbon nanofiber structures through the electropolymerization of pyrrole. Synthetic Metals 157: 282–289 [PMC free article] [PubMed]
- Fletcher BL, McKnight TE, Melechko AV, Simpson ML, Doktycz MJ (2006) Biochemical functionalization of vertically aligned carbon nanofibres. Nanotechnology 17: 2032–2039
- Forster AC, Church GM (2006) Towards synthesis of a minimal cell. Mol Syst Biol 2: 4516924266 [PMC free article] [PubMed]
- Forster S, Antonietti M (1998) Amphiphilic block copolymers in structure-controlled nanomaterial hybrids. Adv Mater 10: 195–217
- Fowlkes JD, Fletcher BL, Hullander ED, Klein KL, Hensley DK, Melechko AV, Simpson ML, Doktycz MJ (2005) Tailored transport through vertically aligned carbon nanofibre membranes; controlled synthesis, modelling, and passive diffusion experiments. Nanotechnology 16: 3101–3109
- Fowlkes JD, Hullander ED, Fletcher BL, Retterer ST, Melechko AV, Hensley DK, Simpson ML, Doktycz MJ (2006a) Molecular transport in a crowded volume created from vertically aligned carbon nanofibres: a fluorescence recovery after photobleaching study. Nanotechnology 17: 5659–5668 [PubMed]
- Fowlkes JD, Melechko AV, Klein KL, Rack PD, Smith DA, Hensley DK, Doktycz MJ, Simpson ML (2006b) Control of catalyst particle crystallographic orientation in vertically aligned carbon nanofiber synthesis. Carbon 44: 1503–1510
- Ghadessy FJ, Ong JL, Holliger P (2001) Directed evolution of polymerase function by compartmentalized self-replication. Proc Natl Acad Sci USA 98: 4552–4557 [PMC free article] [PubMed]
- Gil ES, Hudson SA (2004) Stimuli-reponsive polymers and their bioconjugates. Prog Polym Sci 29: 1173–1222
- Goldberger J, Hochbaum AI, Fan R, Yang PD (2006) Silicon vertically integrated nanowire field effect transistors. Nano Lett 6: 973–977
- Griffiths AD, Tawfik DS (2000) Man-made enzymes—from design to in vitro compartmentalisation. Curr Opin Biotechnol 11: 338–353 [PubMed]
- Griffiths AD, Tawfik DS (2003) Directed evolution of an extremely fast phosphotriesterase by in vitro compartmentalization. EMBO J 22: 24–35 [PMC free article] [PubMed]
- Grosvenor AL, Feltus A, Conover RC, Daunert S, Anderson KW (2000) Development of binding assays in microfabricated picoliter vials: an assay for biotin. Anal Chem 72: 2590–2594 [PubMed]
- Guido NJ, Wang X, Adalsteinsson D, McMillen D, Hasty J, Cantor CR, Elston TC, Collins JJ (2006) A bottom-up approach to gene regulation. Nature 439: 856–860 [PubMed]
- Guo X, Szoka FC (2003) Chemical approaches to triggerable lipid vesicles for drug and gene delivery. Acc Chem Res 36: 335–341 [PubMed]
- Hall D, Minton AP (2003) Macromolecular crowding: qualitative and semiquantitative successes, quantitative challenges. Biochim Biophys Acta-Proteins Proteomics 1649: 127–139 [PubMed]
- Harold FM (2005) Molecules into cells: specifying spatial architecture. Microbiol Mol Biol Rev 69: 544–564 [PMC free article] [PubMed]
- Hasty J, McMillen D, Collins JJ (2002) Engineered gene circuits. Nature 420: 224–230 [PubMed]
- Heller MJ (2002) DNA microarray technology: devices, systems, and applications. Annu Rev Biomed Eng 4: 129–153 [PubMed]
- Hess B, Mikhailov A (1994) Self-organization in living cells. Science 264: 223–224 [PubMed]
- Hess B, Mikhailov A (1995) Microscopic self-organization in living cells: a study of time matching. J Theor Biol 176: 181–184 [PubMed]
- Hildebrand M (2003) Biological processing of nanostructured silica in diatoms. Prog Org Coat 47: 256–266
- Hildebrand M, York E, Kelz JI, Davis AK, Frigeri LG, Allison DP, Doktycz MJ (2006) Nanoscale control of silica morphology and three-dimensional structure during diatom cell wall formation. J Mater Res 21: 2689–2698
- Ho D, Chu B, Lee H, Brooks EK, Kuo K, Montemagno CD (2005) Fabrication of biomolecule-copolymer hybrid nanovesicles as energy conversion systems. Nanotechnology 16: 3120–3132
- Ho RJY, Rouse BT, Huang L (1987) Interactions of target-sensitive immunoliposomes with herpes-simplex virus—the foundation of a sensitive immunoliposome assay for the virus. J Biol Chem 262: 13979–13984 [PubMed]
- Hochachka PW (1999) The metabolic implications of intracellular circulation. Proc Natl Acad Sci USA 96: 12233–12239 [PMC free article] [PubMed]
- Iijima S (1991) Helical microtubules of graphitic carbon. Nature 354: 56–58
- Ishikawa K, Sato K, Shima Y, Urabe I, Yomo T (2004) Expression of a cascading genetic network within liposomes. Febs Lett 576: 387–390 [PubMed]
- Johannessen EA, Weaver JMR, Cobbold PH, Cooper JM (2002) Heat conduction nanocalorimeter for pl-scale single cell measurements. Appl Phys Lett 80: 2029–2031
- Karlsson M, Davidson M, Karlsson R, Karlsson A, Bergenholtz J, Konkoli Z, Jesorka A, Lobovkina T, Hurtig J, Voinova M, Orwar O (2004) Biomimetic nanoscale reactors and networks. Annu Rev Phys Chem 55: 613–649 [PubMed]
- Khandurina J, Guttman A (2002) Microchip-based high-throughput screening analysis of combinatorial libraries. Curr Opin Chem Biol 6: 359–366 [PubMed]
- Kinpara T, Mizuno R, Murakami Y, Kobayashi M, Yamaura S, Hasan Q, Morita Y, Nakano H, Yamane T, Tamiya E (2004) A picoliter chamber array for cell-free protein synthesis. J Biochem 136: 149–154 [PubMed]
- Klein KL, Melechko AV, Rack PD, Fowlkes JD, Meyer HM, Simpson ML (2005) Cu-Ni composition gradient for the catalytic synthesis of vertically aligned carbon nanofibers. Carbon 43: 1857–1863
- Klok HA, Lecommandoux S (2001) Supramolecular materials via block copolymer self-assembly. Adv Mater 13: 1217–1229
- Kong XY, Wang ZL (2003) Spontaneous polarization-induced nanohelixes, nanosprings, and nanorings of piezoelectric nanobelts. Nano Lett 3: 1625–1631
- Kratschmer W, Lamb LD, Fostiropoulos K, Huffman DR (1990) Solid C-60—a new form of carbon. Nature 347: 354–358
- Kroto HW, Heath JR, Obrien SC, Curl RF, Smalley RE (1985) C-60—buckminsterfullerene. Nature 318: 162–163
- Lee YF, Tawfik DS, Griffiths AD (2002) Investigating the target recognition of DNA cytosine-5 methyltransferase HhaI by library selection using in vitro compartmentalisation. Nucleic Acids Res 30: 4937–4944 [PMC free article] [PubMed]
- Long MS, Jones CD, Helfrich MR, Mangeney-Slavin LK, Keating CD (2005) Dynamic microcompartmentation in synthetic cells. Proc Natl Acad Sci USA 102: 5920–5925 [PMC free article] [PubMed]
- Luisi PL, Ferri F, Stano P (2006) Approaches to semi-synthetic minimal cells: a review. Naturwissenschaften 93: 1–13 [PubMed]
- Margulies M, Egholm M, Altman WE, Attiya S, Bader JS, Bemben LA, Berka J, Braverman MS, Chen YJ, Chen ZT, Dewell SB, Du L, Fierro JM, Gomes XV, Godwin BC, He W, Helgesen S, Ho CH, Irzyk GP, Jando SC, Alenquer MLI, Jarvie TP, Jirage KB, Kim JB, Knight JR, Lanza JR, Leamon JH, Lefkowitz SM, Lei M, Li J, Lohman KL, Lu H, Makhijani VB, McDade KE, McKenna MP, Myers EW, Nickerson E, Nobile JR, Plant R, Puc BP, Ronan MT, Roth GT, Sarkis GJ, Simons JF, Simpson JW, Srinivasan M, Tartaro KR, Tomasz A, Vogt KA, Volkmer GA, Wang SH, Wang Y, Weiner MP, Yu PG, Begley RF, Rothberg JM (2005) Genome sequencing in microfabricated high-density picolitre reactors. Nature 437: 376–380 [PMC free article] [PubMed]
- Marijuan PC (1995) Enzymes, artificial cells and the nature of biological information. Biosystems 35: 167–170 [PubMed]
- Mastrobattista E, Taly V, Chanudet E, Treacy P, Kelly BT, Griffiths AD (2005) High-throughput screening of enzyme libraries: In vitro evolution of a beta-galactosidase by fluorescence-activated sorting of double emulsions. Chem Biol 12: 1291–1300 [PubMed]
- McBride J, Treadway J, Feldman LC, Pennycook SJ, Rosenthal SJ (2006) Structural basis for near unity quantum yield core/shell nanostructures. Nano Lett 6: 1496–1501 [PubMed]
- McKnight TE, Peeraphatdit C, Jones SW, Fowlkes JD, Fletcher BL, Klein KL, Melechko AV, Doktycz MJ, Simpson ML (2006) Site-specific biochemical functionalization along the height of vertically aligned carbon nanofiber arrays. Chem Mater 18: 3203–3211
- Mei Q, Fredrickson CK, Jin SG, Fan ZH (2005) Toxin detection by a miniaturized in vitro protein expression array. Anal Chem 77: 5494–5500 [PubMed]
- Melechko AV, Merkulov VI, Lowndes DH, Guillorn MA, Simpson ML (2002) Transition between ‘base’ and ‘tip’ carbon nanofiber growth modes. Chem Phys Lett 356: 527–533
- Melechko AV, Merkulov VI, McKnight TE, Guillorn MA, Klein KL, Lowndes DH, Simpson ML (2005) Vertically aligned carbon nanofibers and related structures: controlled synthesis and directed assembly. J Appl Phys 97: 041301
- Merkulov VI, Hensley DK, Melechko AV, Guillorn MA, Lowndes DH, Simpson ML (2002a) Control mechanisms for the growth of isolated vertically aligned carbon nanofibers. J Phys Chem B 106: 10570–10577
- Merkulov VI, Melechko AV, Guillorn MA, Lowndes DH, Simpson ML (2001) Alignment mechanism of carbon nanofibers produced by plasma-enhanced chemical-vapor deposition. Appl Phys Lett 79: 2970–2972
- Merkulov VI, Melechko AV, Guillorn MA, Lowndes DH, Simpson ML (2002b) Effects of spatial separation on the growth of vertically aligned carbon nanofibers produced by plasma-enhanced chemical vapor deposition. Appl Phys Lett 80: 476–478
- Merkulov VI, Melechko AV, Guillorn MA, Simpson ML, Lowndes DH, Whealton JH, Raridon RJ (2002c) Controlled alignment of carbon nanofibers in a large-scale synthesis process. Appl Phys Lett 80: 4816–4818
- Misteli T (2001) The concept of self-organization in cellular architecture. J Cell Biol 155: 181–185 [PMC free article] [PubMed]
- Monnard PA (2003) Liposome-entrapped polymerases as models for microscale/nanoscale bioreactors. J Membr Biol 191: 87–97 [PubMed]
- Morse DE (1999) Silicon biotechnology: harnessing biological silica production to construct new materials. Trends Biotechnol 17: 230–232
- Murray CB, Norris DJ, Bawendi MG (1993) Synthesis and characterization of nearly monodisperse Cde (E=S, Se, Te) semiconductor nanocrystallites. J Am Chem Soc 115: 8706–8715
- Musyanovych A, Mailander V, Landfester K (2005) Miniemulsion droplets as single molecule nanoreactors for polymerase chain reaction. Biomacromolecules 6: 1824–1828 [PubMed]
- Napoli A, Boerakker MJ, Tirelli N, Nolte RJM, Sommerdijk NAJM, Hubbell JA (2004a) Glucose-oxidase based self-destructing polymeric vesicles. Langmuir 20: 3487–3491 [PubMed]
- Napoli A, Valentini M, Tirelli N, Muller M, Hubbell JA (2004b) Oxidation-responsive polymeric vesicles. Nat Mater 3: 183–189 [PubMed]
- Nardin C, Widmer J, Winterhalter M, Meier W (2001) Amphiphilic block copolymer nanocontainers as bioreactors. EurPhys J E 4: 403–410
- Nguyen-Vu TDB, Chen H, Cassell AM, Andrews R, Meyyappan M, Li J (2006) Vertically aligned carbon nanofiber arrays: an advance toward electrical-neural interfaces. Small 2: 89–94 [PubMed]
- Noireaux V, Libchaber A (2004) A vesicle bioreactor as a step toward an artificial cell assembly. Proc Natl Acad Sci USA 101: 17669–17674 [PMC free article] [PubMed]
- Noireaux V, Bar-Ziv R, Libchaber A (2003) Principles of cell-free genetic circuit assembly. Proc Natl Acad Sci USA 100: 12672–12677 [PMC free article] [PubMed]
- Nojima T, Fujii T, Hosokawa K, Yotsumoto A, Shoji S, Endo I (2000) Cell-free protein synthesis in a microfabricated reactor. Bioprocess Eng 22: 13–17
- Nomura S, Tsumoto K, Hamada T, Akiyoshi K, Nakatani Y, Yoshikawa K (2003) Gene expression within cell-sized lipid vesicles. Chembiochem 4: 1172–1175 [PubMed]
- Oberholzer T, Luisi PL (2002) The use of liposomes for constructing cell models. J Biol Phys 28: 733–744 [PMC free article] [PubMed]
- Park C, Yoon J, Thomas EL (2003) Enabling nanotechnology with self assembled block copolymer patterns. Polymer 44: 6725–6760
- Peng XG, Wickham J, Alivisatos AP (1998) Kinetics of II-VI and III-V colloidal semiconductor nanocrystal growth: ‘Focusing’ of size distributions. J Am Chem Soc 120: 5343–5344
- Petrikovics I, Hong K, Omburo G, Hu QZ, Pei L, McGuinn WD, Sylvester D, Tamulinas C, Papahadjopoulos D, Jaszberenyi JC, Way JL (1999) Antagonism of paraoxon intoxication by recombinant phosphotriesterase encapsulated within sterically stabilized liposomes. Toxicol Appl Pharmacol 156: 56–63 [PubMed]
- Pielak GJ (2005) A model of intracellular organization. Proc Natl Acad Sci USA 102: 5901–5902 [PMC free article] [PubMed]
- Pietrini AV, Luisi PL (2004) Cell-free protein synthesis through solubilisate exchange in water/oil emulsion compartments. Chembiochem 5: 1055–1062 [PubMed]
- Ranquin A, Versees W, Meier W, Steyaert J, Van Gelder P (2005) Therapeutic nanoreactors: combining chemistry and biology in a novel triblock copolymer drug delivery system. Nano Lett 5: 2220–2224 [PubMed]
- Rissin DM, Walt DR (2006) Digital concentration readout of single enzyme molecules using femtoliter arrays and Poisson statistics. Nano Lett 6: 520–523 [PubMed]
- Roco MC (2003) Nanotechnology: convergence with modern biology and medicine. Curr Opin Biotechnol 14: 337–346 [PubMed]
- Rodriguez NM (1993) A Review of Catalytically Grown Carbon Nanofibers. J Mater Res 8: 3233–3250
- Rondelez Y, Tresset G, Tabata KV, Arata H, Fujita H, Takeuchi S, Noji H (2005) Microfabricated arrays of femtoliter chambers allow single molecule enzymology. Nat Biotechnol 23: 361–365 [PubMed]
- Sepp A, Tawfik DS, Griffiths AD (2002) Microbead display by in vitro compartmentalisation: selection for binding using flow cytometry. Febs Lett 532: 455–458 [PubMed]
- Shendure J, Porreca GJ, Reppas NB, Lin XX, McCutcheon JP, Rosenbaum AM, Wang MD, Zhang K, Mitra RD, Church GM (2005) Accurate multiplex polony sequencing of an evolved bacterial genome. Science 309: 1728–1732 [PubMed]
- Shimizu Y, Inoue A, Tomari Y, Suzuki T, Yokogawa T, Nishikawa K, Ueda T (2001) Cell-free translation reconstituted with purified components. Nat Biotechnol 19: 751–755 [PubMed]
- Simpson ML, Sayler GS, Fleming JT, Applegate B (2001) Whole-cell biocomputing. Trends Biotechnol 19: 317–323 [PubMed]
- Smela E (2003) Conjugated polymer actuators for biomedical applications. Adv Mater 15: 481–494
- Spirin AS, Baranov VI, Ryabova LA, Ovodov SY, Alakhov YB (1988) A continuous cell-free translation system capable of producing polypeptides in high yield. Science 242: 1162–1164 [PubMed]
- Sprinzak D, Elowitz MB (2005) Reconstruction of genetic circuits. Nature 438: 443–448 [PubMed]
- Sunami T, Sato K, Matsuura T, Tsukada K, Urabe I, Yomo T (2006) Femtoliter compartment in liposomes for in vitro selection of proteins. Anal Biochem 357: 128–136 [PubMed]
- Tabuchi M, Hino M, Shinohara Y, Baba Y (2002) Cell-free protein synthesis on a microchip. Proteomics 2: 430–435 [PubMed]
- Tawfik DS, Griffiths AD (1998) Man-made cell-like compartments for molecular evolution. Nat Biotechnol 16: 652–656 [PubMed]
- Tong HD, Jansen HV, Gadgil VJ, Bostan CG, Berenschot E, vanRijn CJM, Elwenspoek M (2004) Silicon nitride nanosieve membrane. Nano Lett 4: 283–287
- Vriezema DM, Aragones MC, Elemans JAAW, Cornelissen JJLM, Rowan AE, Nolte RJM (2005) Self-assembled nanoreactors. Chem Rev 105: 1445–1489 [PubMed]
- Wagner RS, Ellis WC (1964) Vapor-liquid-solid mechanism of single crystal growth (new method growth catalysis from impurity whisker epitaxial+large crystals Si E). Appl Phys Lett 4: 89–90
- Walde P, Ichikawa S (2001) Enzymes inside lipid vesicles: preparation, reactivity and applications. Biomol Eng 18: 143–177 [PubMed]
- Wang ZG, Haasch RT, Lee GU (2005) Mesoporous membrane device for asymmetric biosensing. Langmuir 21: 1153–1157 [PubMed]
- Welch GR (1992) An analogical ‘field’ construct in cellular biophysics: history and present status. Prog Biophys Mol Biol 57: 71–128 [PubMed]
- Wolcke J, Ullmann D (2001) Miniaturized HTS technologies—uHTS. Drug Discov Today 6: 637–646 [PubMed]
- Yamamoto T, Nojima T, Fujii T (2002) PDMS-glass hybrid microreactor array with embedded temperature control device. Application to cell-free protein synthesis. Lab on a Chip 2: 197–202 [PubMed]
- Yoshida M, Langer R, Lendlein A, Lahann J (2006) From advanced biomedical coatings to multi-functionalized biomaterials. Polym Rev 46: 347–375
- Yoshimoto M, Wang SQ, Fukunaga K, Walde P, Kuboi R, Nakao K (2003) Preparation and characterization of reactive and stable glucose oxidase-containing liposomes modulated with detergent. Biotechnol Bioeng 81: 695–704 [PubMed]
- Yu W, Sato K, Wakabayashi M, Nakaishi T, Ko-Mitamura EP, Shima Y, Urabe I, Yomo T (2001) Synthesis of functional protein in liposome. J Biosci Bioeng 92: 590–593 [PubMed]
- Zhang L, Melechko AV, Merkulov VI, Guillorn MA, Simpson ML, Lowndes DH, Doktycz MJ (2002) Controlled transport of latex beads through vertically aligned carbon nanofiber membranes. Appl Phys Lett 81: 135–137
Articles from Molecular Systems Biology are provided here courtesy of –
The European Molecular Biology of or relating to a process involving a randomly determined sequence of observations each of which is considered as a sample of one element from a probability distribution.